|
 |
- Search
| Healthc Inform Res > Volume 24(3); 2018 > Article |
Abstract
Objectives
Clinical discharge summaries provide valuable information about patients' clinical history, which is helpful for the realization of intelligent healthcare applications. The documents tend to take the form of separate segments based on temporal or topical information. If a patient's clinical history can be seen as a consecutive sequence of clinical events, then each temporal segment can be seen as a snapshot, providing a certain clinical context at a specific moment. This study aimed to demonstrate a temporal segmentation method of Korean clinical narratives for identifying textual snapshots of patient history as a proof-of-a-concept.
Methods
Our method uses pattern-based segmentation to approximate human recognition of the temporal or topical shifts in clinical documents. We utilized rheumatic patients' discharge summaries and transformed them into sequences of constituent chunks. We built 97 single pattern functions to denote whether a certain chunk has attributes that indicate that it can be a segment boundary. We manually defined the relationships between the pattern functions to resolve multiple pattern matchings and to make a final decision.
The clinical discharge summary in the Electronic Health Record system (EHR) provides detailed descriptions of a patient's clinical events. Physicians describe a patient's disease progress in the document. Because the clinical history tends to be chronologically narrated [1], one of the prominent attributes of clinical documentation is that temporal and causality information comprise the mainstream of writing. In other words, temporal information is embedded in the clinical narrative, and causally related clinical events simultaneously deliver context.
The innate temporal aspect of clinical description makes temporal information processing valuable for utilizing the clinical course described in clinical documents [2]. Thus, temporal processing of clinical data has been a long-standing interest [3]. Indeed, previous clinical natural language processing studies have concerned the extraction of temporal information, such as temporal relation discovery [45678], temporal question answering systems [9], recognition of temporal patterns and visualization of patient's clinical history [101112], as well as temporal segmentation of clinical documents [1314].
Because of their temporality and causality, a sequence of clinical events and related descriptions can be grouped into medical episodes, and the sequence of medical episodes build the temporal structure of a text. A single episode can play the role of a single unit in temporal processing applications. For instance, the causal relationship between clinical entities, which is used in temporal processing applications, can be interpreted within an episode. This study, therefore, attempted to develop a temporal segmentation method for application to clinical narrative documents. A temporal segment is related to a single clinical episode in this study; thus, segmentation can make an intermediate form of the clinical document for clinical temporal processing. Because there is temporal discontinuity [15] between episodes in the terms of text, an important step is to recognize textual cues of the discontinuity in a free text.
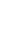
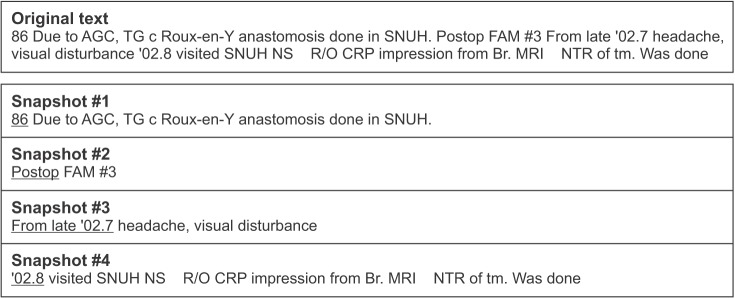
A temporal segment can be interpreted as a snapshot that is literally ŌĆ£a piece of information that delivers readers an idea of what the situation is like at a particular timeŌĆØ (Longman Dictionary of Contemporary English, Pearson, 2009). Temporal segmentation divides a single clinical descriptive text into multiple pieces of narrative texts, and each segment provides temporal coherence between clinical events as shown in Figure 1. From this perspective, temporal segmentation is strongly related to discourse segmentation, as discussed by Allen [16]. A segment is defined as a stretch of clause sequences delivering coherent contents.
The most important indicators of the text structure are ŌĆścue phrasesŌĆÖ. When writing is seen as a linear progression assigning linguistic symbols, authors place signals around the positions leading to a new story. Cue phrases indicate topical or temporal shifts in text structure [1617]. Based on this idea, this study demonstrates a pattern-based segmentation algorithm for clinical narrative texts as a possible way to divide a document into multiple text snippets. Then, this algorithm can make each snippet provide a temporally or topically coherent story for restructuring of the original document. In short, our temporal segmentation algorithm aims to make textual snippets that match the results produced by human readers and that can convey clinical context.
This study used data from the Seoul National University Hospital EHR and was approved by the Institutional Review Board of Seoul National University Hospital (No. 1506-014-677). We obtained 200 discharge summaries of patients hospitalized in the rheumatoid and nephrology departments in 2013 and 2014. We evenly divided the data into training and testing sets, and developed the temporal segmentation algorithm from a portion of the whole training set.
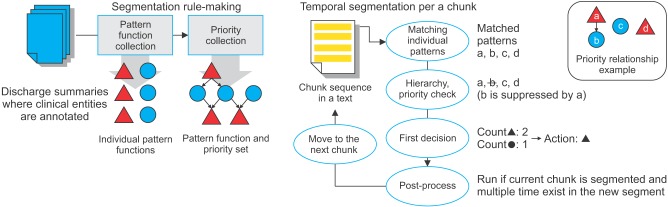
We assume that most segment boundaries exist at the ends of sentences or constituency chunks; thus, it is necessary to preprocess the texts including constituency chunking and sentence boundary detection. The text dataset was manually refined by manual processes. Through constituency chunking, phrases are created from groups of words according to Korean syntactic structure. In addition, texts that convey clinical semantic information must be annotated (i.e., clinical events, temporal anchoring points). The clinical events are related to symptoms, clinical tests, diagnosis, medications, treatments, and clinical department/visit information. Temporal anchoring points are indicated by salient temporal expressions that make a group of temporally coherent textual descriptions related to same temporal information. Then, temporal segmentation using textual cues is performed based on interactions of various segmentation rules. The segmentation logic comprises textual patterns and decision rules for identifying a temporal discontinuity in a document. The sequential steps in the whole process are shown in Figure 2.
As we focused on building the segmentation algorithm, we assumed that both the pre-processing and clinical entity annotation steps had already been developed; therefore, we used clinical texts to which preprocessing had been previously applied and in which the clinical entities had been annotated.
Our segmentation algorithm predicts the positions of segment boundaries. Offsets that present segmentation boundaries are automatically annotated within a text as the output of the algorithm. A segmentation boundary can be any position in general; however, a sentence has phrase constituency that cannot be divided semantically (e.g., a group of a verb and objects in a verb phrase). Thus, our segmentation algorithm assumes that a segmentation boundary exists between chunks in most cases to keep the segmentation outputs rational when a human reads. This process was required to utilize sentence boundary and syntactic chunking information; hence, we manually identified both the sentence and phrase constituency (chunk) boundaries in the corpus.
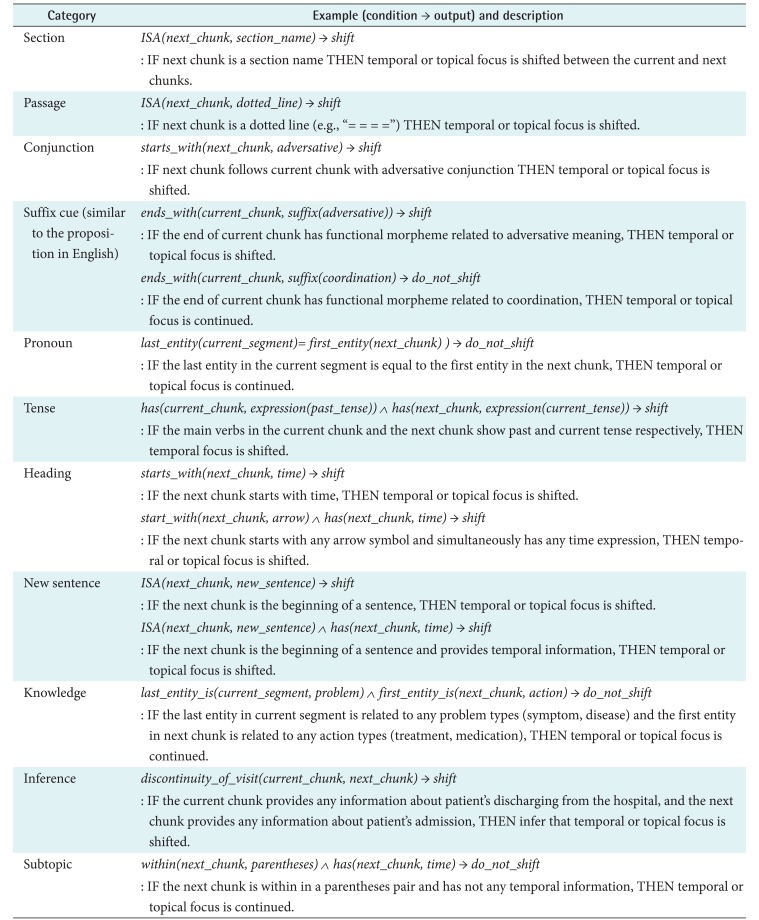
In previous studies, cue phrases, which are used in discourse segmentation, have been defined as linguistic expressions [1618]. Nakhimovsky and Rapaport [15] characterized a cue phrase as a signal that allows a reader to instantly notice a temporal shift when a text is segmented. Probable candidates of cue phrases are generally considered to be pronouns [1619], tense [16], spatial focus [20], and background knowledge [13]. According to previous observations, we found cue phrases for clinical temporal segmentation from our corpora and categorized them into several categories. A cue phrase is called a ŌĆśpattern functionŌĆÖ, and an individual function consists of input, condition, and output values as in a general function. We found 11 categories of pattern functions using the attribute of a chunk as input and providing signals as output whether the chunk boundary should be marked to be shifted or not in the terms of temporality and topic. We reviewed training discharge summaries in sequence, and the pattern functions were collected from the 3,743 chunk boundaries. Each individual pattern function has a condition set. For instance, if we define the ith individual pattern function as pi, and a function pi consists of one or more conditions. The element conditions of pi are grouped by ŌĆśANDŌĆÖ operation, and the individual pattern is matched if and only if an arbitrary input is matched to every condition elements. The output value of each pattern function has one of the segmentation actions in the set {ŌĆśshiftŌĆÖ, ŌĆśdo not shiftŌĆÖ}, and for each element, the temporal or topical focus is shifted at the end of the chunk or not, respectively. Table 1 shows categories and examples of segmentation pattern functions. In total, 97 individual pattern functions are produced in our dataset. Each pattern function is compiled on every chunk, and a pattern marks the action output at the end of a chunk when the pattern is matched to the chunk. If a chunk is not matched to any function, the chunk is marked as ŌĆśshould not be shiftedŌĆÖ.
Because multiple patterns can be matched on a decision point, we use hierarchy information of the pattern functions. We define the ith condition ci and assume two pattern functions pj and pk having same output value. We assume that pj consists of ca and cb, and function pk consists of ca, cb, and cc. In a graphical view, the two pattern functions have a hierarchical relationship when one pattern function is entirely included in another function. By using this concept, the hierarchy relationship is checked among matched functions, and subordinate patterns are rejected in the matched results.
After the hierarchy checking, the segmentation logic is required for the confliction resolution step when two different segmentation actions conflict with each other at one decision point. From our training set, 159 priority relations are collected between conflicted nodes. For example, a single priority relationship between pattern functions pa and pb means that one of them suppresses the other function's output when two functions are simultaneously matched to the same decision point. To avoid sparse data problem, we expand the priority annotation by using a graphical perspective. If pattern pa suppresses pb, and pc is a pattern similar to pa, then the annotation pb also suppresses pc. Finally, the priority checking step suppresses low priority functions, and the logic's final segmentation decision accepts dominated segmentation action. If the final segmentation actions still conflict, then the logic marks the decision point as ŌĆśshould not shiftŌĆÖ. Two or more temporal anchoring points may exist in the segment, although the segmentation logic infers whether a decision point should be segmented; thus, a heuristic post-processing step is applied to resolve the segments. For example, the post-processing first checks whether the temporal anchoring points in the current segment indicate the same time point or not, and it performs extra segmentation within the segment by using some heuristic rules. Figure 3 graphically illustrates the detailed segmentation process.
The patterns and the priority relations were incrementally collected from the training data. We observed segmentation patterns for each document, and the pattern function collection was updated when new pattern functions appeared. As a consequence, half of the pattern functions were discovered from the first 5 documents, and the frequency of the newly discovered patterns from each new document tended to be very low after the 5 documents. In other words, collected segmentation patterns in the previous documents can properly segment most of the following documents. According to this tendency, the pattern collection was iterated for exploring 50 documents, and the patterns were arranged by testing the patterns on another development set of 15 documents.
The human judge group consisted of two medical doctors and one biomedical researcher. Also, they were native Korean speakers who could use English fluently. The human judges were provided a Web-based interface for the temporal segmentation evaluation. The interface presented the algorithm's segmentation outputs. Through the interface, the judges were asked whether they agreed or disagreed with the algorithm's single predictions at each segmentation boundary where the algorithm predicted a boundary. In addition, they provided corrections at each segment if they did not agree with the algorithm's prediction. The corrections were used for reference in the quantitative evaluation.
The prediction results produced by the temporal segmentation algorithm were assessed in comparison to multiple human experts' agreement on the segmentation output. During the evaluation, three human judges were independently asked whether they agreed with each segmentation output and to make corrections of the segmentation boundaries. Using the individual experts' corrections as reference segmentations, we evaluated our model in terms of precision, recall, and F1 for each segmentation boundary. The test dataset for temporal segmentation comprised 1,243 clinical sentences and 1,849 chunks in 30 clinical documents (average number of sentences per document, 41.4). We noted that we only used a certain portion out of the whole test set to make the human experts' revision process tolerable with a proper number of documents to evaluate.
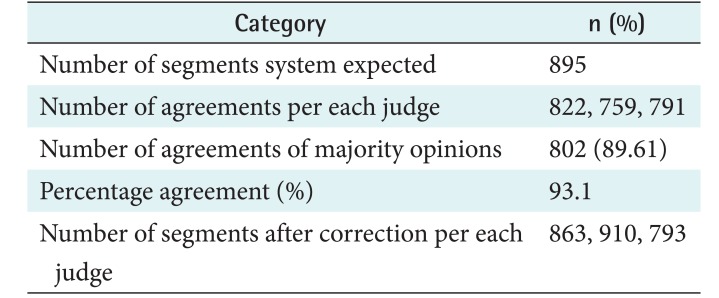
The segmentation algorithm scanned the given 1,849 chunks and made 895 individual temporal segments in the test set. The human judges marked their opinion whether they agreed or disagreed at each decision position and made corrections if necessary. The number of times the human judges agreed with the algorithm results out of the 895 temporal segments were 822, 759, and 791 per judge, respectively. The majority opinion at each decision point was used as the final decision. That is, if two or all judges agreed at a certain point, then the final decision was marked as ŌĆśagreedŌĆÖ; otherwise, it was marked as ŌĆśdisagreedŌĆÖ. The majority opinion agreed with the algorithm's results for 802 decision points out of the 895 points (89.61%). Inter-rater agreement was calculated according to [2122]. The percentage agreement is used to calculate the inter-rater agreement among multiple judges; it is the ratio of the number of agreements with the major opinion over the number of possible agreements with the major opinion. In our evaluation, the number of possible agreements was 2,685 (= 895 ├Ś 3), and the human judges agreed with the majority opinion 2,501 times. Consequently, the agreement percentage was 93.1%. As previously stated, each judge provided corrections of the algorithm's outputs during the evaluation, and they made 863, 910, and 793 segments, respectively. Table 2 summarizes the qualitative evaluation results.
By using the judges' segmentation correction as human segmentation references independently, the algorithm's outputs were quantitatively measured in terms of precision, recall, and F1-score. For measuring segmentation outputs, precision is the number of correct boundary predictions over the number of the algorithm's predictions; recall is the number of correct predictions over the number of segmentation boundaries in the judge's reference. F1-score is the harmonic mean of the precision and recall. Table 3 presents the evaluation results. The first row shows the averaged value for the references made by the three independent human judges. The other rows show the quantitative evaluation results for the references made by individual judges.
The reason we used the qualitative evaluation as the first measurement was that judges' temporal granularity inconsistently varies depending on the human cognition if explicit temporal information is absent. When we asked the judges to make segmentation boundary corrections, their corrections tended to differ from each other. For instance, judge #2 preferred to make fine-grained segments, resulting in 910 segments, whereas judge #3 preferred to make relatively coarse-grained segments, resulting 793 segments. This means multiple forms of snapshots are allowed. We were concerned that the segmentation boundaries predicted by the algorithm with which humans may agree could be identified as incorrect if the reference decision boundary was pre-defined and fixed before testing of the algorithm. Thus, on our evaluation the algorithm's predictions were given to the human judges, and the judges determined whether or not they agreed. In addition, building a single consented segmentation reference was challenging. A general method for creating a single reference annotation is to use majority opinion; however, when the judges' corrections were merged, we observed that marginal errors could make the textual snippet awkward. For this reason, each individual's corrections were tested independently in the quantitative evaluation.
Document or discourse segmentation algorithms are generally evaluated by Pk [23] and WindowDiff [24], allowing near-misses; however, in our temporal segmentation exact segmentation boundary prediction is preferred because human readers recognize linguistic awkwardness if subtle segmentation boundary errors occur. Thus, our quantitative evaluation for the segmentation algorithm uses measurements that only accept exactly correct boundary predictions.
Beyond simple lexical cues, segmentation signals from domain knowledge were exploited; however, our rules could not cover signals requiring more intelligent sense, such as a sense of clinical location. Some location information without temporal information can signal readers that a temporal shift has occurred. For instance, ŌĆśadmissionŌĆÖ and ŌĆśfollow-upŌĆÖ events may conflict in the terms of temporal information. Another example concerns distinguishing clinical events during a patient's hospitalization from those during an outpatient visit following hospitalization. For instance, two events, a routine treatment with a high-dose immunosuppressant during a patient's hospitalization and the next routine during an outpatient visit should be distinguished even if the explicit temporal information is absent. Although some human judges recognize the temporal shift between two events, it is difficult to translate into rules.
Our motivation was to build a method to providing intermediate forms for clinical temporal processing applications utilizing temporal snapshots. If a temporal normalization method [25] is applied to temporal segments to chronologically arrange the snapshots, the temporal structure information would be helpful in creating a timeline visualization of a patient's history and mining semantic relationships, such as the temporal order of clinical events and causal relationships [1026272829]. However, there are some points that should be considered to improve our knowledge further. First, summarizing an idea into a kind of linguistic presentation seems too complex to be abstracted without any information loss. As well, clinical records contain a significant number of arbitrary tabular layouts of words or nested structures of temporal information in terms of syntax. Our method is a linear progression of a text; thus, the method seems to have difficulty, especially for the arbitrary layout structure beyond the linear structure. These issues are challenges that must be addressed for the building of further temporal processing applications.
This paper presented a temporal segmentation method for capturing snapshots of patient histories in Korean clinical discharge summaries. Each segment provides a temporally or topically coherent story for restructuring the original document. Human judges were asked whether they agreed with the temporal segmentation results, and the percentage of agreement with the majority opinion was 89.61%. Temporal segmentation of clinical free texts has not been fully explored in the medical informatics domain, and only a few related studies have been previously conducted [1330]. Although this study has the limitation that the algorithm relied on human intervention for its construction, this study provided an important opportunity to advance the understanding of clinical document segmentation regarding the temporal coherence of clinical events. This study demonstrated a trial implementing the temporal processing of clinical texts based on intuitive segmentation features. We plan to improve our method by adding machine learning approaches that minimize human intervention in this process in the future. This would lead to more generalizable temporal segmentation methods for clinical narrative documents.
Acknowledgments
This work was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (No. NRF-2015R1D1A1A01058075), and also supported by a grant of the Korea Health Technology R&D Project through the Korea Health Industry Development Institute, funded by the Ministry of Health & Welfare, Republic of Korea (No. HI14C1277).
References
1. Zhou L, Friedman C, Parsons S, Hripcsak G. System architecture for temporal information extraction, representation and reasoning in clinical narrative reports. AMIA Annu Symp Proc 2005;2005:869-873.



2. Combi C, Shahar Y. Temporal reasoning and temporal data maintenance in medicine: issues and challenges. Comput Biol Med 1997;27(5):353-368. PMID: 9397339.


3. Madkour M, Benhaddou D, Tao C. Temporal data representation, normalization, extraction, and reasoning: A review from clinical domain. Comput Methods Programs Biomed 2016;128:52-68. PMID: 27040831.



4. Verhagen M, Gaizauskas R, Schilder F, Hepple M, Katz G, Pustejovsky J. SemEval-2007 task 15: TempEval temporal relation identification 4th International Workshop on Semantic Evaluations; 2007 Jun 23ŌĆō24. Prague, Czech Republic; p. 75-80.

5. Bramsen PJ. Doing time: inducing temporal graphs [dissertation]. Cambridge (MA): Massachusetts Institute of Technology; 2006.

6. Zhou L, Parsons S, Hripcsak G. The evaluation of a temporal reasoning system in processing clinical discharge summaries. J Am Med Inform Assoc 2008;15(1):99-106. PMID: 17947618.



7. Pustejovsky J, Stubbs A. Increasing informativeness in temporal annotation 5th Linguistic Annotation Workshop; 2011 Jun 23ŌĆō24. Portland, OR; p. 152-160.

8. Dligach D, Miller T, Lin C, Bethard S, Savova G. Neural temporal relation extraction 15th Conference of the European Chapter of the Association for Computational Linguistics (Volume 2, Short Papers); 2017 Apr 3ŌĆō7. Valencia, Spain; p. 746-751.

9. Tao C, Solbrig HR, Sharma DK, Wei WQ, Savova GK, Chute CG. Time-oriented question answering from clinical narratives using semantic-web techniques. In: Patel-Schneider PF, Pan Y, Hitzler P, Mika P, Zhang L, Pan JZ, Horrocks I, editors. International Semantic Web Conference. Heidelberg: Springer; 2010. p. 241-256.

10. Park H, Choi J. V-Model: a new perspective for EHR-based phenotyping. BMC Med Inform Decis Mak 2014;14:90. PMID: 25341558.



11. Jung H, Allen J, Blaylock N, De Beaumont W, Galescu L, Swift M. Building timelines from narrative clinical records: initial results based-on deep natural language understanding BioNLP 2011 Workshop; 2011 Jun 23ŌĆō24. Portland, OR; p. 146-154.

12. Monroe M, Lan R, Lee H, Plaisant C, Shneiderman B. Temporal event sequence simplification. IEEE Trans Vis Comput Graph 2013;19(12):2227-2236. PMID: 24051789.


13. Angelova G, Boytcheva S. Towards temporal segmentation of patient history in discharge letters Workshop on Biomedical Natural Language Processing; 2011 Sep 15ŌĆō16. Hissar, Bulgaria; p. 49-54.

14. Bramsen P, Deshpande P, Lee YK, Barzilay R. Finding temporal order in discharge summaries. AMIA Annu Symp Proc 2006;2006:81-85.



15. Nakhimovsky A, Rapaport WJ. Discontinuities in narratives 12th Conference on Computational Linguistics; 1988 Aug 22ŌĆō27. Budapest, Hungary; p. 465-470.

17. Nakhimovsky A. Aspect, aspectual class, and the temporal structure of narrative. Comput Linguist 1988;14(2):29-43.

18. Grosz BJ, Sidner CL. Attention, intentions, and the structure of discourse. Comput Linguist 1986;12(3):175-204.

19. Anderson A, Garrod SC, Sanford AJ. The accessibility of pronominal antecedents as a function of episode shifts in narrative text. Q J Exp Psychol A 1983;35(3):427-440.

20. Maybury MT. Using discourse focus, temporal focus, and spatial focus to generate multisentential text 5th International Workshop on Natural Language Generation; 1990 Jun 3ŌĆō6. Dawson, PA; p. 70-78.

21. Passonneau RJ, Litman DJ. Empirical analysis of three dimensions of spoken discourse: segmentation, coherence, and linguistic devices. In: Hovy EH, Scott DR, editors. Computational and conversational discourse. Heidelberg: Springer; 1996. p. 161-194.

22. Marcu D. The theory and practice of discourse parsing and summarization. Cambridge (MA): MIT Press; 2000.

23. Beeferman D, Berger A, Lafferty J. Text segmentation using exponential models 2nd Conference on Empirical Methods in Natural Language Processing; 1997 Aug 1ŌĆō2. Providence, RI; p. 37-46.

24. Pevzner L, Hearst MA. A critique and improvement of an evaluation metric for text segmentation. Comput Linguist 2002;28(1):19-36.

25. Kim Y, Choi J. Recognizing temporal information in korean clinical narratives through text normalization. Healthc Inform Res 2011;17(3):150-155. PMID: 22084809.



26. Chapman WW, Nadkarni PM, Hirschman L, D'Avolio LW, Savova GK, Uzuner O. Overcoming barriers to NLP for clinical text: the role of shared tasks and the need for additional creative solutions. J Am Med Inform Assoc 2011;18(5):540-543. PMID: 21846785.



27. Seol JW, Yi W, Choi J, Lee KS. Causality patterns and machine learning for the extraction of problem-action relations in discharge summaries. Int J Med Inform 2017;98:1-12. PMID: 28034407.


28. Uzuner O, South BR, Shen S, DuVall SL. 2010 i2b2/VA challenge on concepts, assertions, and relations in clinical text. J Am Med Inform Assoc 2011;18(5):552-556. PMID: 21685143.



29. Sun W, Rumshisky A, Uzuner O. Evaluating temporal relations in clinical text: 2012 i2b2 Challenge. J Am Med Inform Assoc 2013;20(5):806-813. PMID: 23564629.