|
 |
- Search
| Healthc Inform Res > Volume 22(3); 2016 > Article |
Abstract
Objectives
The number of deaths caused by cardiovascular disease and stroke is predicted to reach 23.3 million in 2030. As a contribution to support prevention of this phenomenon, this paper proposes a mining model using a naïve Bayes classifier that could detect cardiovascular disease and identify its risk level for adults.
Methods
The process of designing the method began by identifying the knowledge related to the cardiovascular disease profile and the level of cardiovascular disease risk factors for adults based on the medical record, and designing a mining technique model using a naïve Bayes classifier. Evaluation of this research employed two methods: accuracy, sensitivity, and specificity calculation as well as an evaluation session with cardiologists and internists. The characteristics of cardiovascular disease are identified by its primary risk factors. Those factors are diabetes mellitus, the level of lipids in the blood, coronary artery function, and kidney function. Class labels were assigned according to the values of these factors: risk level 1, risk level 2 and risk level 3.
Results
The evaluation of the classifier performance (accuracy, sensitivity, and specificity) in this research showed that the proposed model predicted the class label of tuples correctly (above 80%). More than eighty percent of respondents (including cardiologists and internists) who participated in the evaluation session agree till strongly agreed that this research followed medical procedures and that the result can support medical analysis related to cardiovascular disease.
Cardiovascular disease is deadly and can attack adults at any time [12]. It has become a major health problem in both developed and developing countries, and it is cited as the number one cause of death throughout the world each year. In 2008, around 17.3 million deaths were caused by cardiovascular disease. More than 3 million of these deaths occur in people under 60 years old. The number of deaths caused by cardiovascular disease accounts for around 4% of total deaths in high-income countries and around 42% in low-income countries. The number of deaths caused by cardiovascular disease and stroke is predicted to reach 23.3 million in 2030 [3]. The World Health Organization has noted that 60% of deaths in adults were caused by coronary artery disease in 2014 [4]. In Indonesia, the number of people with the cardiovascular disease increases each year. In some cases, this condition causes disability and social-economic problem for the patient, his/her family, community, and even country. In 2013, based on doctor diagnoses, the prevalence of cardiovascular disease in Indonesia is around 0.5% or approximately 883,447 people, while based on symptoms, it is thought to be around 1.5%, or approximately around 2.6 million people [5].
Since the number of patients with cardiovascular disease is increasing, early detection of this disease needed. Cardiovascular disease can be detected through the biochemical testing (blood, urine or tissue testing) of samples obtained from a patient. Other indicators are basic biochemical risk factors for heart disease detection, namely, blood pressure, cigarette smoking, glucose, cholesterol (Chol), low-density-lipoprotein cholesterol (LDL-C), high-density-lipoprotein cholesterol (HDL-C), and physical inactivity. Furthermore, biochemical testing of the blood is used to detect cardiovascular disease. Such testing detects fats, cholesterol, and lipid components of blood, including LDL, HDL, triglycerides, blood sugar, and glycosylated hemoglobin, which is measured for diabetes detection. In addition, C-reactive protein (CRP) and others protein markers, such as poly-protein A1 and B are used to detect inflammation that might lead to cardiovascular disease [6789]. Thus the number of deaths of patients with cardiovascular disease can be reduced through accurate diagnosis through biochemical tests and appropriate treatment.
There is a huge amount of data available within the healthcare industry [10]. All of this data is stored in huge databases of electronic medical records systems [1112]. Drowning in data but starving for knowledge, and the need to provide cardiovascular disease detection with accurate diagnosis has become an enormous challenge for hospitals [413141516]. Data can be explored for evaluation purposes [10], but in fact, hospitals have not explored them appropriately yet. A lot of hidden information from the data has not been mined yet [21718] and the discovery of hidden patterns from data is rarely exploited [19]. The mining of medical records for analysis purposes could be developed to guide and support the clinical decision-making process [142021].
Some recent research on data mining for healthcare has utilized medical records, especially for cardiovascular disease, and various methods and predictors have been applied as seen in Table 1. Also, data mining techniques have been conducted with various comparable methods, such as naïve Bayes, decision tree, neural network [22], and classification [23] or clustering. Comparisons have shown that naïve Bayes has the highest accuracy among many approaches [14202425]. A naïve (or simple) Bayesian classifier based on Bayes' theorem is a probabilistic statistical classifier; the term "naïve" indicates conditional independence among features or attributes [26]. All predictor attributes are identified from interview sessions, and then used in a naïve Bayes classifier to produce valuable knowledge for medical analysis purposes. Archana and Elangovan [27] analyzed the advantages and disadvantages of several data mining techniques. The results showed that naïve Bayes requires a short computational time and achieves good performance, but naïve Bayes requires a very large number of records to obtain good results. A naïve Bayesian classifier is more accurate than other classifiers [28], as reported by Subbalakshmi et al. [29]. Other kinds of medical data obtained from electrocardiography (ECG), echocardiography, or coronary angiography are known as evidence for cardiovascular diagnosis. Mining signal and image data is still an open research area.
This research used medical records, and we propose a data mining model using a naïve Bayes classifier, which can detect cardiovascular disease and identify its risk level for adults. The selected predictor attributes are mostly based on the basic biochemical attributes related to cardiovascular disease, and patients are categorized into two groups (normal and cardiovascular risk) based on blood and urine testing. Subsequently, for more detail analysis, the cardiovascular risk group is divided into three risk levels, namely, risk levels 1, 2, and 3.
Techniques such as fact finding (interview), technical analysis and evaluation were applied in the research. Open-ended questions were designed for the interviews, and cardiologists and internists at a private hospital in the Southern part of Jakarta participated. One data source for this research was the blood chemical laboratory of Mayapada Hospital, which stores the results of patients' blood and urine tests. Such testing is mandatory to examine liver and kidney health, check for diabetes, fat accumulation, and cardiac function, and to examine the metabolism and organs function. Thus, this testing can examine the potential of cardiovascular disease. The head nurse in the catheterization laboratory provided information related to characteristics, types, and level of risk factors for cardiovascular disease for adults. The information consisted of causes of cardiovascular disease, diagnosis of cardiovascular disease at the catheterization laboratory, and cardiovascular risk factor.
Another data source collected from cardiac risk assessment was used to indicate the level of risk factors. Finally, the results of interview sessions were used to identify predictor attributes for each group of cardiovascular risk factors and to determine the class labels related to cardiovascular disease risk level.
A total of 60,589 records and 38 medical attributes were obtained from this database. They consisted of medical checkup result and blood chemistry test results. They then passed into three processes: data reduction, data cleaning, and data generalization. A stepwise backward elimination method was used to select predictor attributes, and the attributes not selected were eliminated. This process is called data reduction. Subsequently, the selected attributes were used as predictor attributes for analysis. The selected predictor attributes were the following: birthdate, sex, glucose level (GLU, GLU2J), cholesterol level (CHOL), triglyceride level (TRIG), HDL and LDL level, UREA, creatine level (CREA), uric acid (UA), creatine kinase level (CK, CKMB), lactate level (LDH), and troponin level (TROPK, TROPT). The second process was data cleaning. This process identified incomplete, incorrect, inaccurate, and irrelevant parts of the data (called dirty data) and then replaced, modified or deleted them. After the data cleaning process, only 50,528 records remained. For mining process consideration, related to the mining process, since almost all of the attributes in this research were numeric-type data (integer or real value), while sex is binary-type data, sex was excluded in this research. Although sex was not considered in this research, the diagnosis result was valid. This result was validated by a cardiologist and internal medicine specialist at Mayapada Hospital.
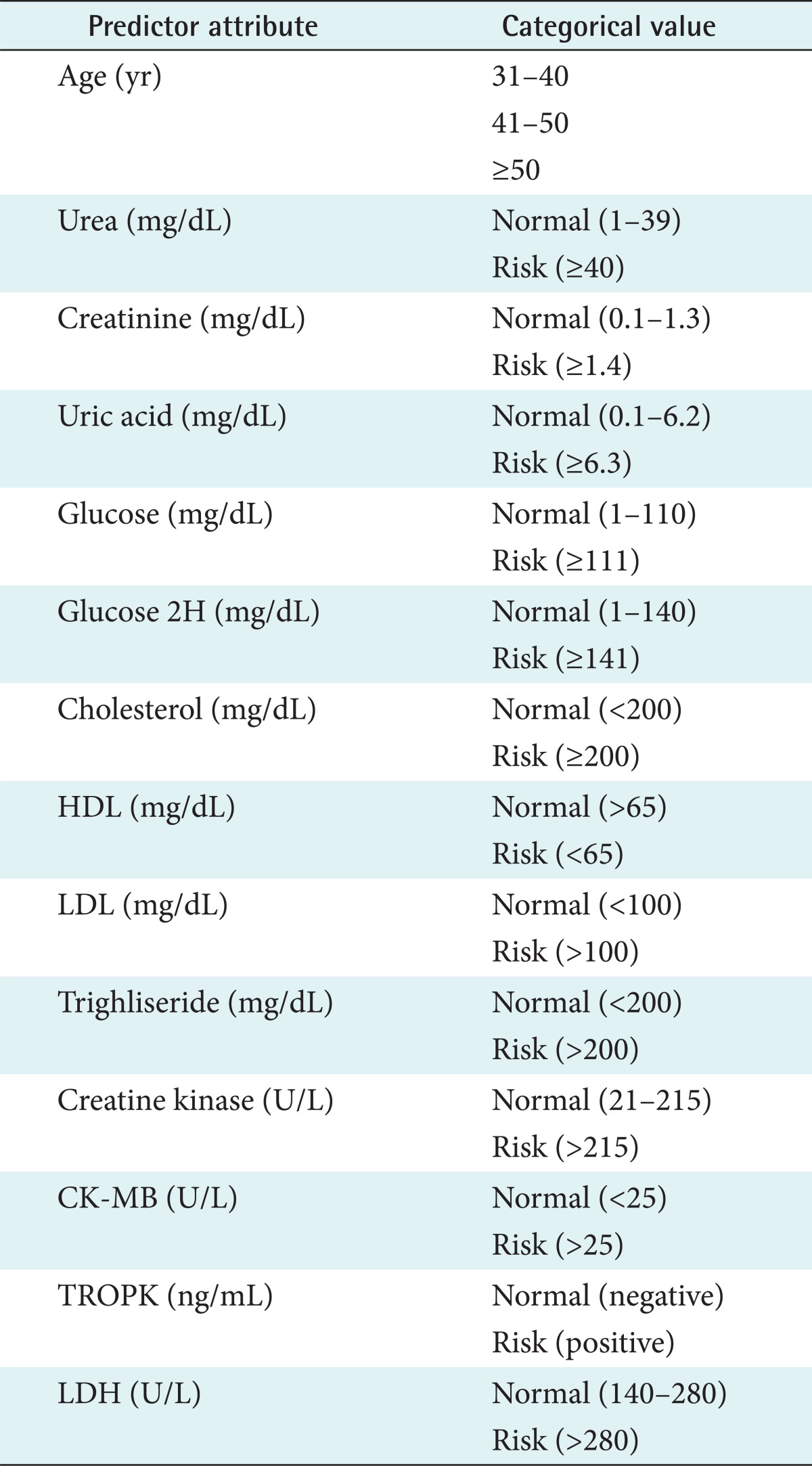
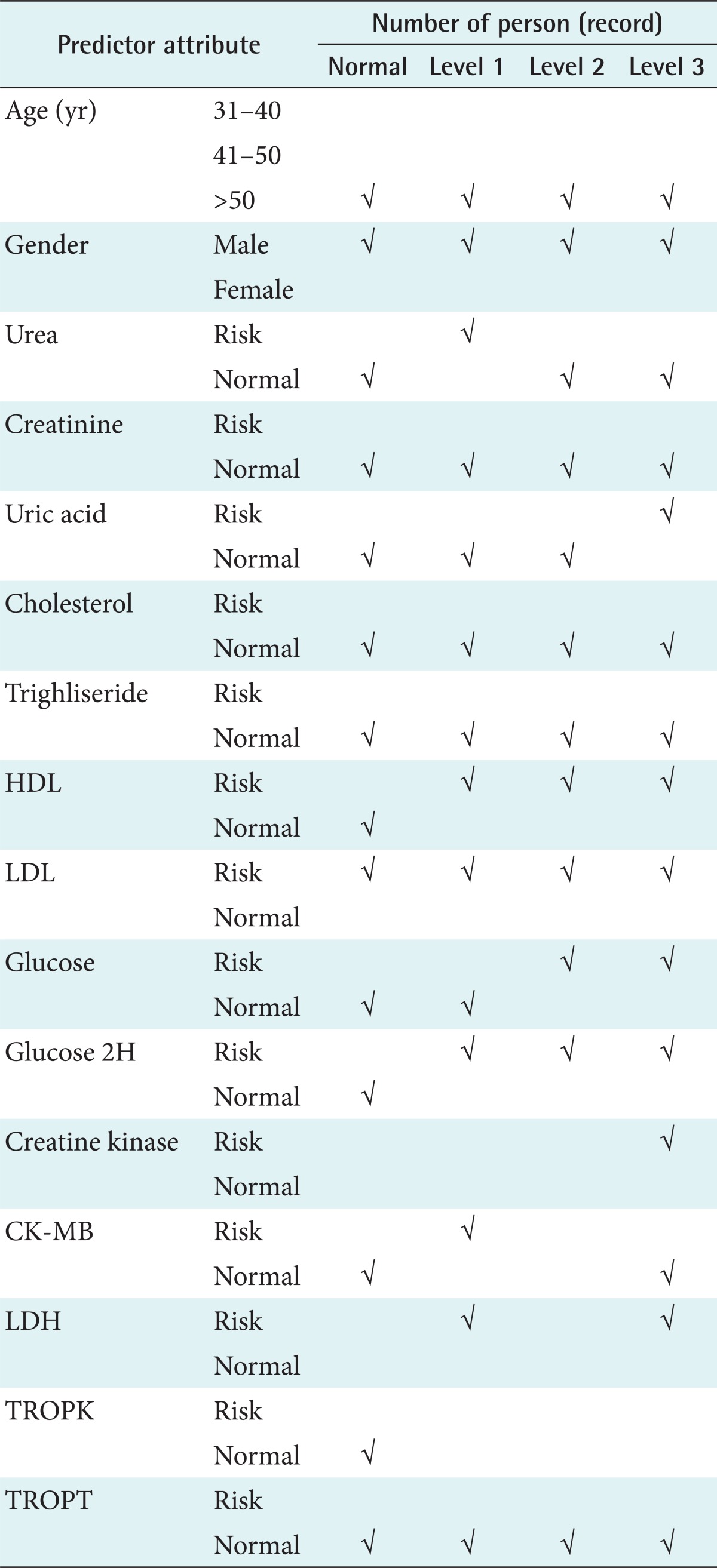
The third process was data generalization. This process changed low-level data (e.g., numeric values for an attribute age) into high-level data through the conversion of data values into categorical data (e.g., young, middle-aged, and senior) or by reducing the number of dimensions to summarize data into a concept space involving fewer dimensions. Table 2 showed all predictor attributes and their categorical values. All categorical values were determined based on medical laboratory testing standards.
The results of the interview sessions were used to address the following questions: What are the characteristics of cardiovascular disease in adults? What were the risk factors? The answers to those questions were then used to identify the primary factors of cardiovascular disease. The primary factors of cardiovascular disease were identified as the following: diabetes mellitus (attributes: Glucose, Glucose 2H), the level of lipids in blood (attributes: Cholesterol, HDL, LDL, TRIG), coronary artery function (attributes: CK, CKMB, TROPK, LDH), and kidney function (attributes: UREA, CREA, UA). Thus, those attributes have been identified as the predictor attributes as well. Furthermore, an analysis based on the relationship of each primary factor to cardiovascular system was conducted. Subsequently, based on the analysis results, any reference values that were considered abnormal (beyond normal standard on medical laboratory test) were determined, and the class labels related to cardiovascular disease risk level were determined.
The class labels consist of three categories:
Risk level 1: At least one attribute of one of the main factors of cardiovascular disease (lipid, diabetes, coronary artery function or kidney function) was above the normal standard.
Risk level 2: Two predictor attributes were above the normal standard for at least two primary factors, and each attribute of the primary factors has at least one attribute above normal standard.
Risk level 3: Predictor attributes were above the normal standard for three primary factors (lipid, diabetes, and kidney function) and each attribute of primary factors has at least one attribute above normal standard and the coronary artery function included in this analysis.
In this section, we present in greater detail how the naïve Bayes classifier is used to detect cardiovascular disease and identify its risk level. The naïve Bayes classifier, or simple Bayes classifier, consists of two main components, namely, a training set of tuples and their associated class label. Blood and urine test results from the clinical laboratory database were used as a training dataset, while class labels were defined based on the results of the interview sessions.
A likelihood value was calculated by comparing the observed distribution among classes of tuples covered by a rule with the expected distribution that would result if the rule made predictions at random. A likelihood value describes the probability of the observed data generated from the model conditioned on the given parameter (normal, risk level 1, risk level 2, or risk level 3). Subsequently, the prior value is calculated by summarizing all class targets divided by the number of records. The prior value for each class is shown in Table 3.
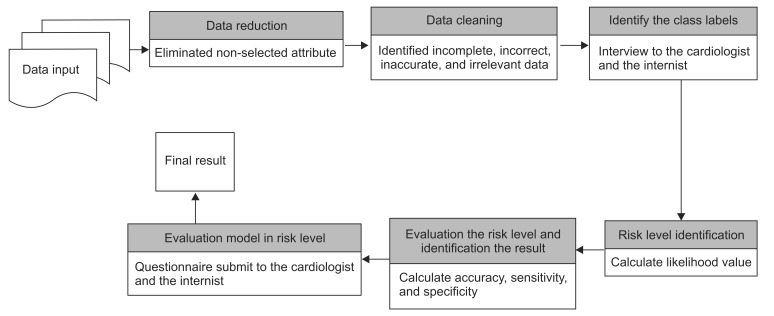
Afterward, the posterior value was calculated. This value is obtained by multiplication of the likelihood by the prior value. The posterior probability of a classification can be defined as "What is the probability that a particular object belongs to a class which was given its observed feature values?" If the posterior result for the normal class is greater than the posterior value for level 1, 2 or 3, then the model will classify the data into the normal level and vice versa; if the posterior values for level 1, level 2, and level 3 are greater than the other comparison posterior value then the model will classify the data into the risk level which is used as the target level. Finally, all steps of naïve Bayes risk level modeling are shown in Figure 1.
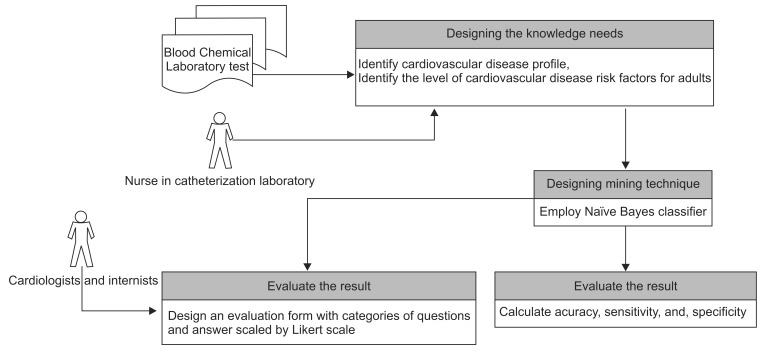
Evaluation sessions were conducted in the same private hospital with cardiologists, an internist, and the head nurse of the catheterization laboratory. Four categories of questions were designed to be scaled by five-level opinions (Likert scale), namely, strongly agree, agree, neither, disagree, and strongly disagree, based on the hospital's medical procedures. The resulting application has benefits for doctors and other medical personnel to support medical analysis related to cardiovascular disease with the same level of accuracy or accuracy very similar to that achieved when manually conducted by a cardiologist or internist, especially for adults. All of the steps of the research on this method are shown in Figure 2.
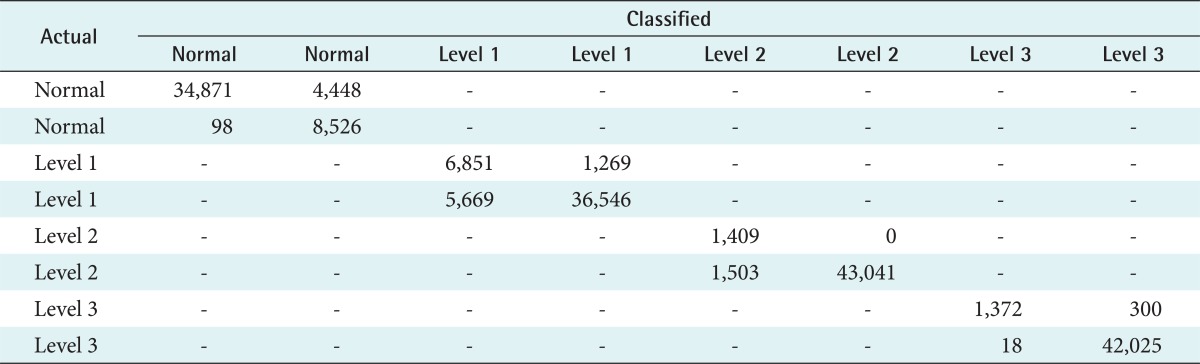
A naïve Bayes evaluation model was formed by testing data and class target (risk level status). Evaluation results were generated by calculation of accuracy level, sensitivity, specificity, and error rate from testing the model using the data. Evaluation of the naïve Bayes classification model calculated true positive, false positive, false negative, and true negative rates for each class target namely: normal level, risk level 1, risk level 2, and risk level 3. This evaluation is summarized in the confusion matrix shown in Table 4.
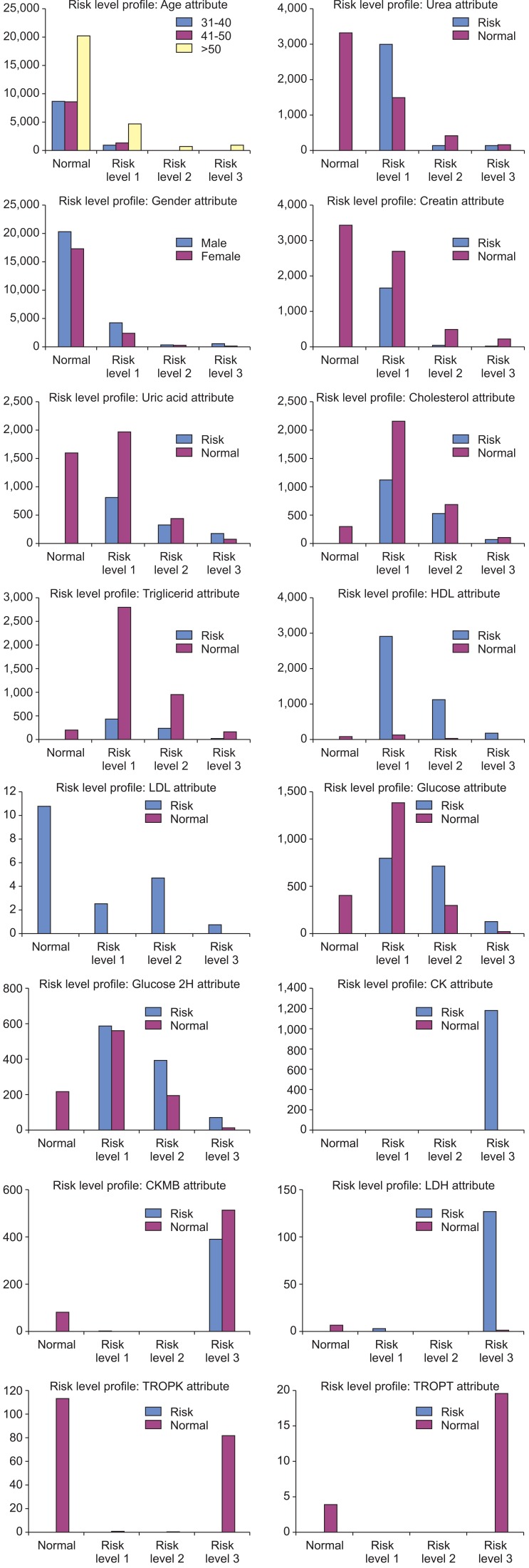
The class label results for the whole dataset and the predictor attributes are shown in Figure 3.
The characteristics for each level of risk of cardiovascular disease can be described as seen in Table 5. The dominant risk factors for cardiovascular disease were identified as Urea, UA, HDL, LDL, glucose, CK, CKMB, and LDH.
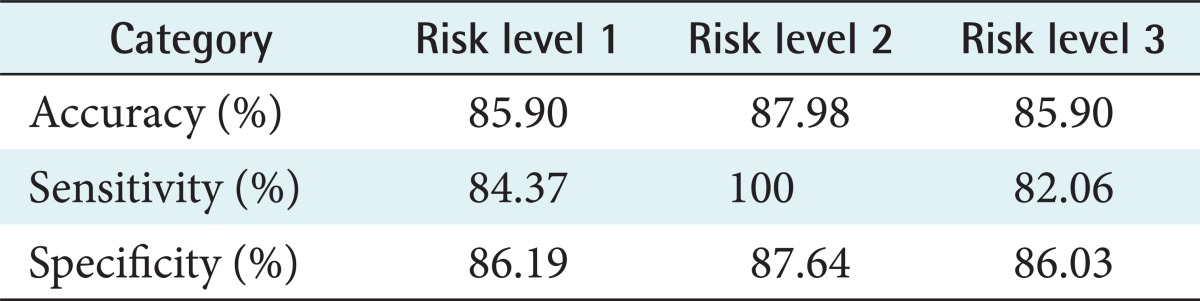
Finally, the accuracy and the benefits of the model were tested and evaluated using two methods: (1) calculation of accuracy, sensitivity and specificity values as conducted by Wiharto et al. [30] and (2) evaluation of the model through an interview evaluation session. The results of the first evaluation method are shown in Table 6.
The sensitivity value (true positive recognition rate) shows the ratio of the positive tuples that were correctly labeled by the classifier to positive tuples. The sensitivity values for each level were 84.37%, 100%, and 82.06%, respectively. The specificity value (true negative recognition rate) shows the ratio of the negative tuples that were correctly labeled by the classifier to negative tuples. The specificity values for each level were 86.19%, 87.64%, and 86.03%, respectively. The accuracy value shows the ratio of correctly classified samples to the total number of tests samples. The accuracy values for each level were 85.90%, 87.98%, and 85.90%, respectively.
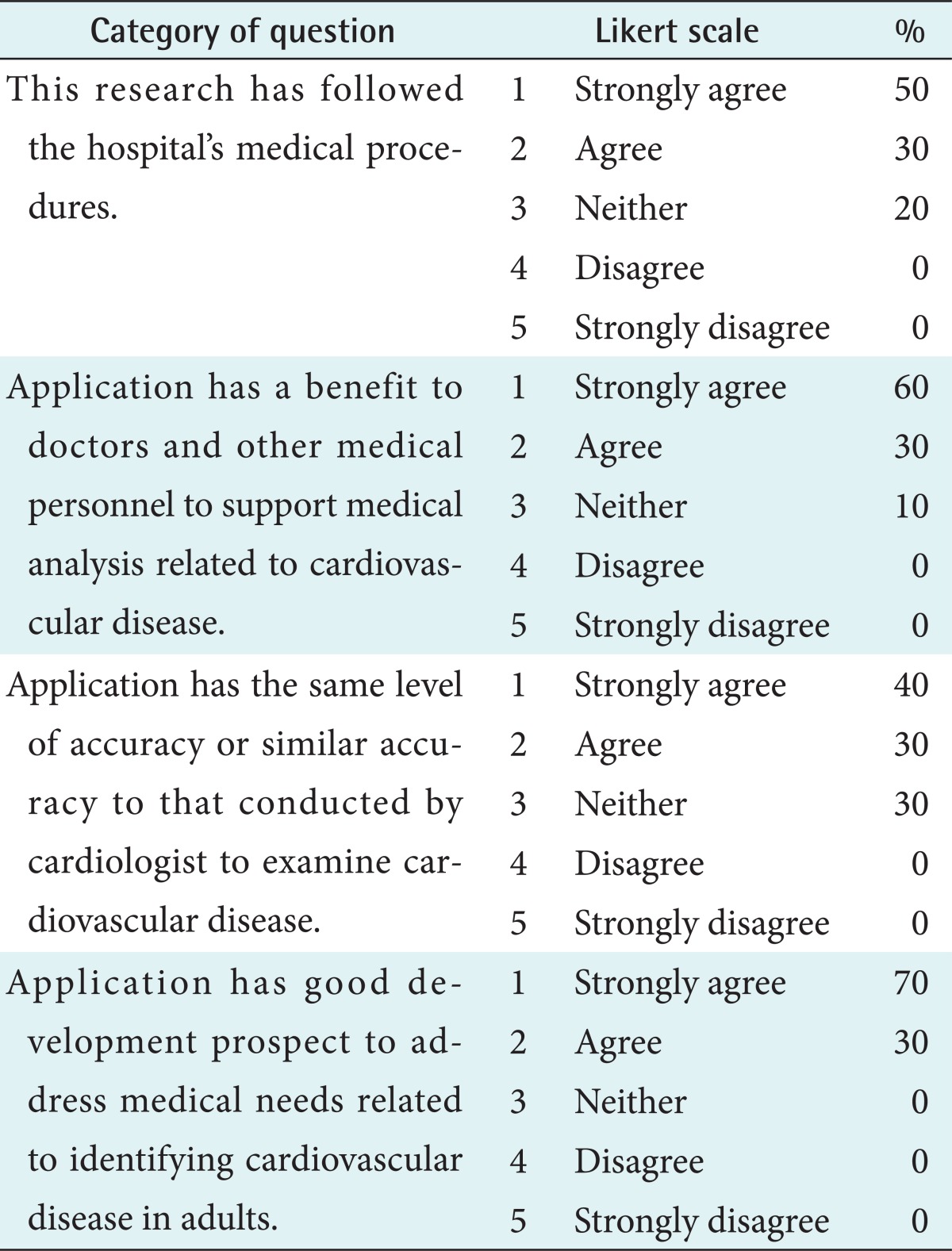
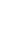
An evaluation session was conducted as the final step to evaluate the benefit of cardiovascular disease risk level for adults using the naïve Bayes classifier result. This session was carried out through four categories of questions administered to cardiologists and an internist at the hospital, and the evaluation session results are shown in Table 7.
More than eighty percent of respondents agree till strongly agreed that this research followed medical procedures and that the model has benefit to doctors and can support medical analysis related to cardiovascular disease with very similar accuracy to the analysis that would be conducted by a cardiologist.
A naïve Bayes approach to heart disease detection was employed by previous researchers such as Soni et al. [20]. They employed a naïve Bayes approach for heart disease prediction with an accuracy value of 86.53% using 22 predictor attributes: sex, age, chest pain, fasting blood sugar, resting electrographic results, exercise induced angina, the slope of the peak exercise ST segment, number of major vessels colored by fluoroscopy, blood pressure, serum cholesterol and maximum heart rate achieved. This research used two biochemical attributes namely blood sugar and cholesterol.
Evaluation of this research was carried out by two methods. First, accuracy, sensitivity, and specificity were calculated, and each value was above 80% for all risk levels. Second, the model was assessed through an evaluation session with cardiologists and an internist. More than 80% of respondents (including cardiologists and internists) who participated in the evaluation session agree till strongly agreed that this research followed medical procedures and that the result can support medical analysis related to cardiovascular disease.
A data mining model was developed with a clinical laboratory database using a naïve Bayes classifier to detect cardiovascular risk, and it was tested for its accuracy in predicting three levels of risk.
The proposed model was trained and validate against data testing. Measurement of accuracy, sensitivity, and specificity showed that this model has an accuracy level greater than eighty percent to detect cardiovascular disease at the each three risk level, especially for adults. seventy percent of respondents (including cardiologists and internists) in the evaluation session strongly agreed this model has the contribution in medical science to support medical analysis and detection related to cardiovascular disease.
Other data associated with cardiovascular disease analysis, such as ECG, echocardiography or coronary angiography can be used for further research. In addition, other classification techniques, such as decision tree, rule-based classification, and many others classification techniques can be conducted and compared to find the most valid one.
Acknowledgments
We gratefully acknowledge the support of institutions in the implementation of this research. Many thanks to the Mayapada hospital, Jakarta, Indonesia, which supported the availability of data and medical personnel for testing and data validation. We also thank the Research and Technology Transfer Office of Bina Nusantara University, which provided internal funding for the research until the publication of this article.
References
1. Shouman M, Turner T, Stocker R. Using data mining techniques in heart disease diagnosis and treatment. Proceedings of Japan-Egypt Conference on Electronics Communications and Computers (JEC-ECC); 2012 Mar 6-9. Alexandria; Egypt: p. 173-177.

2. Ishtake SH, Sanap SA. Intelligent heart disease prediction system using data mining techniques. Int J Healthc Biomed Res 2013;1(3):94-101.

3. Ministry of Health Republic of Indonesia. Healthy environment, healthy heart [Internet]. Jakarta: Ministry of Health Republic of Indonesia; 2014. cited at 2016 Jan 17. Available from: http://www.depkes.go.id/article/view/201410080002/lingkungan-sehat-jantung-sehat.html

4. World Health Organization, Department of Health Statistics and Information Systems. Global Health Estimates: key figures and tables [Internet]. Geneva: World Health Organization; 2016. cited at 2016 Jan 17. Available from: http://www.who.int/healthinfo/global_burden_disease/en

5. Ministry of Health Republic of Indonesia. Heart health situation [Internet]. Jakarta: Ministry of Health Republic of Indonesia; 2014. cited at 2016 Jan 17. Available from: http://www.depkes.go.id/folder/view/01/structure-publikasi-pusdatin-info-datin.html

6. Marcovina SM, Crea F, Davignon J, Kaski JC, Koenig W, Landmesser U, et al. Biochemical and bioimaging markers for risk assessment and diagnosis in major cardiovascular diseases: a road to integration of complementary diagnostic tools. J Intern Med 2007;261(3):214-234. PMID: 17305644.


7. Miao C, Bao M, Xing A, Chen S, Wu Y, Cai J, et al. Cardiovascular health score and the risk of cardiovascular diseases. PLoS One 2015;10(7):e0131537PMID: 26154254.



8. National Institutes of Health, National Heart, Lung, and Blood Institute. How is heart disease diagnosed? [Internet]. Washington (DC): National Institutes of Health; 2014. cited at 2016 Jan 17. Available from: http://www.nhlbi.nih.gov/health/health-topics/topics/hdw/diagnosis

9. Sun X, Jia Z. A brief review of biomarkers for preventing and treating cardiovascular diseases. J Cardiovasc Dis Res 2012;3(4):251-254. PMID: 23233766.



10. El-Sappagh SH, El-Masri S, Riad AM, Elmogy M. Data mining and knowledge discovery: applications, techniques, challenges and process models in healthcare. Int J Eng Res Appl 2013;3(3):900-906.

11. Cortes PL, Cortes EG. Hospital information systems: a study of electronic patient records. J Inf Syst Technol Manag 2011;8(1):131-154.

12. Qiu Y, Zhen S, Zhou M, Li L. Continuously improve the medical care quality and hospital management level through medical information system construction. J Transl Med 2012;10(Suppl 2):A56.


13. Butler J, Kalogeropoulos A. Hospital strategies to reduce heart failure readmissions: where is the evidence? J Am Coll Cardiol 2012;60(7):615-617. PMID: 22818073.


14. Anbarasi M, Anupriya E, Iyengar NC. Enhanced prediction of heart disease with feature subset selection using genetic algorithm. Int J Eng Sci Technol 2010;2(10):5370-5376.

15. Nishimura RA, Otto CM, Bonow RO, Carabello BA, Erwin JP 3rd, Guyton RA, et al. 2014 AHA/ACC guideline for the management of patients with valvular heart disease: executive summary: a report of the American College of Cardiology/American Heart Association Task Force on Practice Guidelines. J Am Coll Cardiol 2014;63(22):2438-2488. PMID: 24603192.


16. Krum H, Driscoll A. Management of heart failure. Med J Aust 2013;199(5):334-339. PMID: 23992190.


17. Holzinger A, Dehmer M, Jurisica I. Knowledge discovery and interactive data mining in bioinformatics: state-of-the-art, future challenges and research directions. BMC Bioinformatics 2014;15(Suppl 6):I1PMID: 25078282.

18. Roque FS, Jensen PB, Schmock H, Dalgaard M, Andreatta M, Hansen T, et al. Using electronic patient records to discover disease correlations and stratify patient cohorts. PLoS Comput Biol 2011;7(8):e1002141PMID: 21901084.



19. Palaniappan S, Awang R. Intelligent heart disease prediction system using data mining techniques. Int J Comput Sci Netw Secur 2008;8(8):343-350.

20. Soni J, Ansari U, Sharma D, Soni S. Predictive data mining for medical diagnosis: an overview of heart disease prediction. Int J Comput Appl 2011;17(8):43-48.

21. Sakthimurugan T, Poonkuzhali S. An effective retrieval of medical records using data mining techniques. Int J Pharm Sci Health Care 2012;2(2):72-78.

22. Chaurasia V, Pal S. Early prediction of heart diseases using data mining techniques. Caribb J Sci Technol 2013;1:208-217.

23. Das R, Turkoglu I, Sengur A. Effective diagnosis of heart disease through neural networks ensembles. Expert Syst Appl 2009;36(4):7675-7680.

24. Sudhakar K, Manimekalai M. Propose a enhanced framework for prediction of heart disease. Int J Eng Res Appl 2015;5(4):1-6.

25. Rajkumar A, Reena GS. Diagnosis of heart disease using datamining algorithm. Glob J Comput Sci Technol 2010;10(10):38-43.

26. Jabbar MA, Deekshatulu BL, Chandra P. Classification of heart disease using k-nearest neighbor and genetic algorithm. Procedia Technol 2013;10:85-94.

27. Archana S, Elangovan K. Survey of classification techniques in data mining. Int J Comput Sci Mob Appl 2014;2(2):65-71.

28. Han J, Kamber M. Data mining: concepts and techniques. 3rd ed. Amsterdam: Morgan Kaufmann; 2012.

29. Subbalakshmi G, Ramesh K, Rao MC. Decision support in heart disease prediction system using naive Bayes. Indian J Comput Sci Eng 2011;2(2):170-176.

30. Wiharto W, Kusnanto H, Herianto H. Intelligence system for diagnosis level of coronary heart disease with K-Star algorithm. Healthc Inform Res 2016;22(1):30-38. PMID: 26893948.